HerpesDRG - Herpesvirus Drug Resistance Genotyping
HSV1, HSV2, HCMV, VZV & HHV6b
Upload a file to begin processing
Clinical overview - mutations > 10% frequency
Identified resistant mutants
Graphics showing location of identified resistance mutations, in context to all database mutations
This page desciribes usage and results from HerpesDRG
Inputs
First select the virus references you want to use in processing from the drop-down list.
- FASTA (Whole genome, or fragments i.e. UL54 & UL89
- Varaint Call Format >= Ver4.0
- Varscan tabluated format
Clinical Overview
- High level (any mutation confers a fold change above 15)
- Moderate level (maximal fold change returned is between 5-15)
- Low resistance (maximal fold change returned is between 2-5)
- No Resistance (maximal fold change returned was less than 2, or only anecdotal 'Polymorphism' was returned)
- Resistant, magnitude unknown ( only anecdotal resistant data was returned)
- NA (no mutations returned)

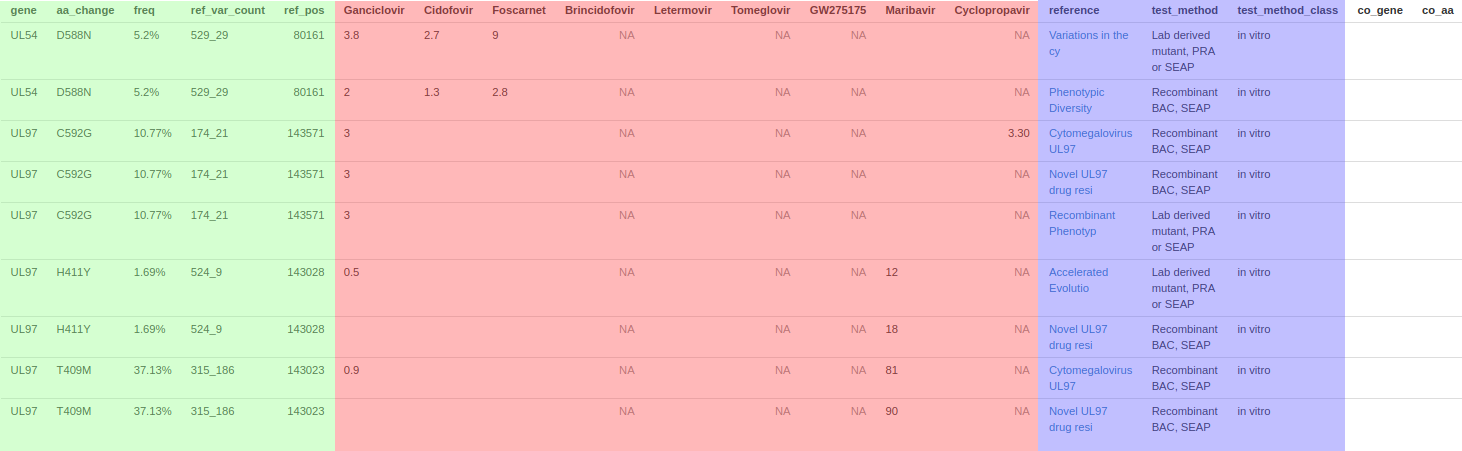
Identified resistant mutants
green: Genetic information. Gene is Amino-acid change, freq is percentage of variant_reads / (all reads), ref_var_count is absolute values, ref_pos is relative nucleotide position to virus ncbi reference.
red: Antiviral sensitivity change. Each header is an antiviral drug recorded within the database, a single mutation can map to many entries & an entry can map to many drugs. Values are the EC50 fold change determined relative to a wild type virus. 'Resistant' or 'Polymorphism' in entries are typically associated with anecdotal or less defined entries & users should consult the references
blue: Reference information. reference is a hyperlink to the original reference where available. test_method is a brief summary of the method used to derive the entry as descriped in the references article, test_method_class in even more brief.
co_gene and co_aa: some database entries contain tested viruses with co-occuring marker transferred mutations. Where flagged these columns are filled.
Disclaimer
We clearly state here that the tool is not validated for use in a diagnostic or clinical care context.
We clearly state here that any file uploaded should be devoid of patient identifiable features, such as filename.
When you upload a file to use this web service, you are accepting these terms.
Privacy Policy
Every time you access the hosted webserver at a UCL web address we reserve the right to register the following data for statistical and troubleshooting purposes.
- Date and time of access
- IP adress.
The contents of uploaded files are only temporarily stored, and not read manually.
last updated: Nov 2023
The effective use of antivirals is crucial to the management of herpesvirus infections in immuno-compromised hosts. Genotypic resistance testing is a widely adopted part of this strategy as resistance has been identified for all licensed drugs. HerpesDRG is the first open knowledgebase linking all published herpes mutations to their in-vitro impact on antiviral sensitivity. This webserver is a simple tool to enable resistance genotyping from common virus sequence data.
Database
The tabular database has been manually curated by reading published literature and extracting key information. Each entry is a mutation containing the link between virus, gene, AAchange & EC50 fold change relative to a control. The data comes from both marker transfer in-vitro experiments and characterised clinical isolates. We update the database manually circa every quarter with new data, you can track updates in GitHub (see below).
Method:
Open Source
Anyone is welcome to propose changes to the knowledgebase or R package. i.e. If you spot mistakes or omissions in the either contact oscar.charles.18@ucl.ac.uk or make a pull request.
Link to the herpesdrg-db github reposisitory
Link to the herpesdrg R package github repository
Recent News
2026 Feb - We had an issue with the webserver causing a blackout for 2 weeks.
Contact & Referencing
The main developer is Oscar Charles who can be contacted by starting an issue in GitHub here. To reference HerpesDRG please cite this article.
AckowledgementsWe would like to thank the MRC and Wellcome Trust as sponsors of this work.


